New substance library to accelerate the search for active compounds
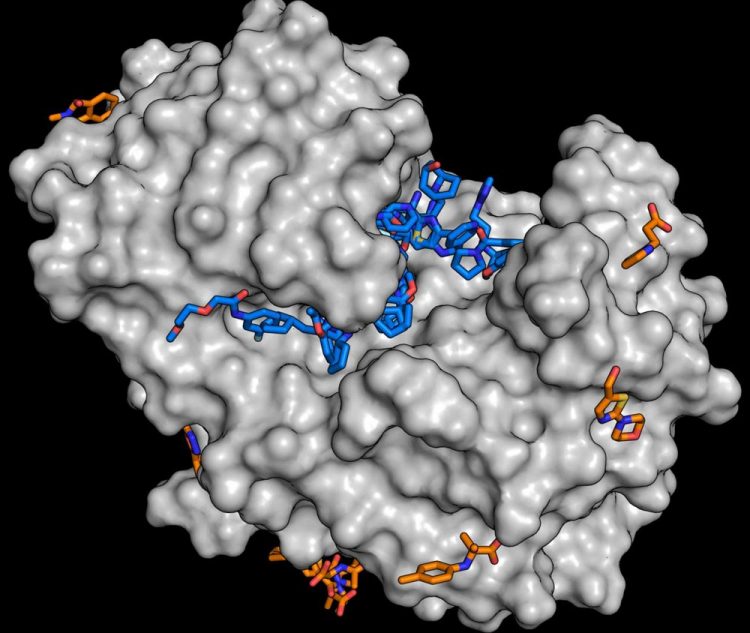
For the study, the enzyme endothiapepsin (grey) was combined with molecules from the fragment library. The analysis shows that numerous substances are able to dock to the enzyme (blue and orange molecules). Every substance found is a potential starting point for the development of larger molecules. Credit: J. Wollenhaupt/HZB
For several years now, the team of the Macromolecular Crystallography Department (MX) at HZB headed by Dr. Manfred Weiss together with the Drug Design Group headed by Prof. Gerhard Klebe (University of Marburg) has therefore been working on building up what are known as fragment libraries.
These consist of small organic molecules (fragments) with which the functionally important cavities on the surface of proteins can be probed and mapped. Protein crystals are saturated with the fragments and then analysed using powerful X-ray light. This allows three-dimensional structural information to be obtained at levels of atomic resolution.
Among other things, it is possible to find out how well a specific molecule fragment docks to the target protein. The development of these substance libraries took place as part of the joint Frag4Lead research project and was funded by the German Federal Ministry of Education and Research (BMBF).
The MX team (MX stands for Macromolecular Crystallography) has now published the design of a chemically diverse fragment library called the “F2X-Universal” library, which consists of 1,103 compounds.
A representative selection of 96 compounds has been extracted from this library, which is referred to as the F2X Entry Screen. In the course of publishing the library, this selection has now been successfully tested and validated by the MX team of the HZB at the MAX IV X-ray source in Lund, Sweden and at BESSY II.
In the study, the HZB and MAX IV teams verified the efficiency of the F2X Entry library by screening endothiapepsin and the Aar2/RnaseH protein complex as the target enzymes. In the next step, the MX team will use the entire universal library.
“For the current study, the fragment screening experts at HZB – BESSY II worked very closely with the FragMAX project team at MAX IV”, said Dr. Uwe Müller from the MX team at HZB who helped to set up the three MX beamlines at BESSY II as well as the BioMAX beamline at MAX IV.
“This enabled both partners to further develop their own technology platforms and use them for imaging the functional surfaces of different proteins. This will be an excellent basis for future collaboration between MAX IV and HZB.”
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Pinpointing hydrogen isotopes in titanium hydride nanofilms
Although it is the smallest and lightest atom, hydrogen can have a big impact by infiltrating other materials and affecting their properties, such as superconductivity and metal-insulator-transitions. Now, researchers from…

A new way of entangling light and sound
For a wide variety of emerging quantum technologies, such as secure quantum communications and quantum computing, quantum entanglement is a prerequisite. Scientists at the Max-Planck-Institute for the Science of Light…

Telescope for NASA’s Roman Mission complete, delivered to Goddard
NASA’s Nancy Grace Roman Space Telescope is one giant step closer to unlocking the mysteries of the universe. The mission has now received its final major delivery: the Optical Telescope…



