Artificial intelligence successfully predicts protein interactions
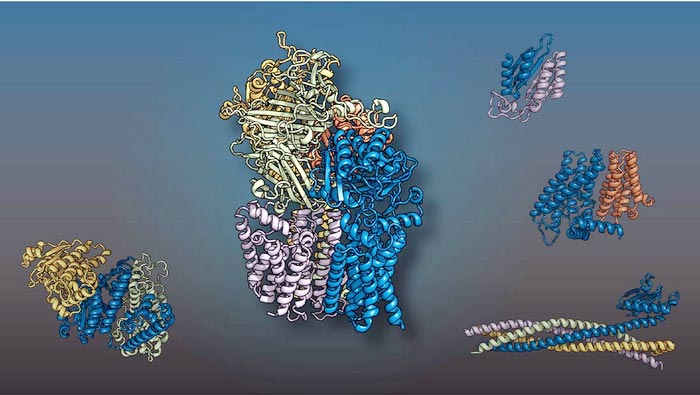
The yeast proteins shown in different colors come together as two-, three-, four-, and five-member complexes like 3D puzzle pieces to execute cellular functions. An international team led by researchers at UT Southwestern and the University of Washington predicted the structures using artificial intelligence techniques.
Credit: UT Southwestern Medical Center
Research led by UT Southwestern and the University of Washington could lead to a wealth of drug targets.
UT Southwestern and University of Washington researchers led an international team that used artificial intelligence (AI) and evolutionary analysis to produce 3D models of eukaryotic protein interactions. The study, published in Science, identified more than 100 probable protein complexes for the first time and provided structural models for more than 700 previously uncharacterized ones. Insights into the ways pairs or groups of proteins fit together to carry out cellular processes could lead to a wealth of new drug targets.
“Our results represent a significant advance in the new era in structural biology in which computation plays a fundamental role,” said Qian Cong, Ph.D., Assistant Professor in the Eugene McDermott Center for Human Growth and Development with a secondary appointment in Biophysics.
Dr. Cong led the study with David Baker, Ph.D., Professor of Biochemistry and Dr. Cong’s postdoctoral mentor at the University of Washington prior to her recruitment to UT Southwestern. The study has four co-lead authors, including UT Southwestern Computational Biologist Jimin Pei, Ph.D.
Proteins often operate in pairs or groups known as complexes to accomplish every task needed to keep an organism alive, Dr. Cong explained. While some of these interactions are well studied, many remain a mystery. Constructing comprehensive interactomes – or descriptions of the complete set of molecular interactions in a cell – would shed light on many fundamental aspects of biology and give researchers a new starting point on developing drugs that encourage or discourage these interactions. Dr. Cong works in the emerging field of interactomics, which combines bioinformatics and biology.
Until recently, a major barrier for constructing an interactome was uncertainty over the structures of many proteins, a problem scientists have been trying to solve for half a century. In 2020 and 2021, a company called DeepMind and Dr. Baker’s lab independently released two AI technologies called AlphaFold (AF) and RoseTTAFold (RF) that use different strategies to predict protein structures based on the sequences of the genes that produce them.
In the current study, Dr. Cong, Dr. Baker, and their colleagues expanded on those AI structure-prediction tools by modeling many yeast protein complexes. Yeast is a common model organism for fundamental biological studies. To find proteins that were likely to interact, the scientists first searched the genomes of related fungi for genes that acquired mutations in a linked fashion. They then used the two AI technologies to determine whether these proteins could be fit together in 3D structures.
Their work identified 1,505 probable protein complexes. Of these, 699 had already been structurally characterized, verifying the utility of their method. However, there was only limited experimental data supporting 700 of the predicted interactions, and another 106 had never been described.
To better understand these poorly characterized or unknown complexes, the University of Washington and UT Southwestern teams worked with colleagues around the world who were already studying these or similar proteins. By combining the 3D models the scientists in the current study had generated with information from collaborators, the teams were able to gain new insights into protein complexes involved in maintenance and processing of genetic information, cellular construction and transport systems, metabolism, DNA repair, and other areas. They also identified roles for proteins whose functions were previously unknown based on their newly identified interactions with other well-characterized proteins.
“The work described in our new paper sets the stage for similar studies of the human interactome and could eventually help in developing new treatments for human disease,” Dr. Cong added.
Dr. Cong noted that the predicted protein complex structures generated in this study are available to download from ModelArchive. These structures and others generated using this technology in future studies will be a rich source of research questions for years to come, she said.
Dr. Cong is a Southwestern Medical Foundation Scholar in Biomedical Research. Other UTSW researchers who contributed to this study include Jing Zhang and Josep Rizo, Ph.D., who holds the Virginia Lazenby O’Hara Chair in Biochemistry.
Collaborating institutions include: Harvard University, Wayne State University, Cornell University, MRC Laboratory of Molecular Biology, Memorial Sloan Kettering Cancer Center, Gerstner Sloan Kettering Graduate School of Biomedical Sciences, Fred Hutchinson Cancer Research Center, Columbia University, University of Würzburg in Germany, St Jude Children’s Research Hospital, FIRC Institute of Molecular Oncology in Milan, Italy, and the National Research Council, Institute of Molecular Genetics in Rome, Italy.
This work was supported by Southwestern Medical Foundation, the Cancer Prevention and Research Institute of Texas (CPRIT) (RP210041), Amgen, Microsoft, the Washington Research Foundation, Howard Hughes Medical Institute, National Science Foundation (DBI 1937533), National Institutes of Health (R35GM118026, R01CA221858, R35GM136258, R21AI156595), UK Medical Research Council (MRC_UP_1201/10), HHMI Gilliam Fellowship, the Deutsche Forschungsgemeinschaft (KI-562/11-1, KI-562/7-1), AIRC investigator and the European Research Council Consolidator (IG23710 and 682190), Defense Threat Reduction Agency (HDTRA1-21-1-0007), and the National Energy Research Scientific Computing Center.
About UT Southwestern Medical Center
UT Southwestern, one of the nation’s premier academic medical centers, integrates pioneering biomedical research with exceptional clinical care and education. The institution’s faculty has received six Nobel Prizes, and includes 25 members of the National Academy of Sciences, 16 members of the National Academy of Medicine, and 14 Howard Hughes Medical Institute Investigators. The full-time faculty of more than 2,800 is responsible for groundbreaking medical advances and is committed to translating science-driven research quickly to new clinical treatments. UT Southwestern physicians provide care in about 80 specialties to more than 117,000 hospitalized patients, more than 360,000 emergency room cases, and oversee nearly 3 million outpatient visits a year.
Media Contact
UT Southwestern Medical Center
UT Southwestern Medical Center
news@utsouthwestern.edu
Office: 214-648-3404
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Innovative 3D printed scaffolds offer new hope for bone healing
Researchers at the Institute for Bioengineering of Catalonia have developed novel 3D printed PLA-CaP scaffolds that promote blood vessel formation, ensuring better healing and regeneration of bone tissue. Bone is…

The surprising role of gut infection in Alzheimer’s disease
ASU- and Banner Alzheimer’s Institute-led study implicates link between a common virus and the disease, which travels from the gut to the brain and may be a target for antiviral…

Molecular gardening: New enzymes discovered for protein modification pruning
How deubiquitinases USP53 and USP54 cleave long polyubiquitin chains and how the former is linked to liver disease in children. Deubiquitinases (DUBs) are enzymes used by cells to trim protein…



